Description
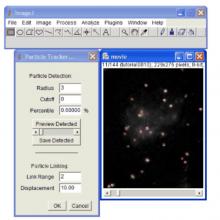
Easy-to-use, computationally efficient, two- and three-dimensional, feature point-tracking tool for the automated detection and analysis of particle trajectories as recorded by video imaging in cell biology.
The tracking process requires no apriori mathematical modelling of the motion, it is self-initialising, it discriminates spurious detections, and it can handle temporary occlusion as well as particle appearance and disappearance from the image region.
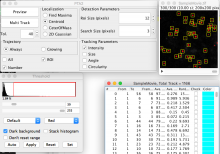
The plugin is well suited for video imaging in cell biology relying on low-intensity fluorescence microscopy. It allows the user to visualize and analyze the detected particles and found trajectories in various ways:
- Preview and save detected particles for separate analysis
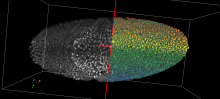
- Global non progressive view on all trajectories
- Focused progressive view on individually selected trajectory
- Focused progressive view on trajectories in an area of interest
It also allows the user to find trajectories from uploaded particles position and information text files and then to plot particles parameters vs. time - along a trajectory