Data Science in Cell Imaging (DSCI) course material
Graduate course at the department of Software and Information Systems Engineering at Ben-Gurion University of the Negev.
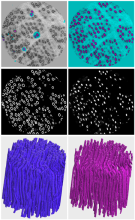
The recent explosion in high-content, dynamic and multidimensional imaging data is transforming cell imaging into a “Data Science” field. This course will review the state-of-the-art in visualizing, processing, integrating and mining massive cell image data sets, deciphering complex patterns and turning them into new biological insight. It will include a mix of approaches in computer science, machine learning and computer vision applied to bio-imaging data.