Introduction to 3D Analysis with 3D ImageJ Suite
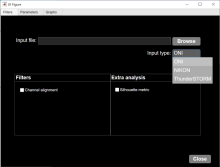
The 3D ImageJ Suite is a set of algorithms and tools (mostly ImageJ plugins) developed since 2010, originally for 3D analysis of fluorescence microscopy. Since then, the plugins have been widely used and cited more than 200 times in biological journals. In this presentation we will give a general introduction to the tools available in the 3D ImageJ Suite : filtering, 3D segmentation for spots and nuclei, and 3D analysis. A graphical interface to manage 3D objects, the 3DManager, was also developed and will be presented.