MosaicSuite
Type
Requires
Implementation Type
Supported image dimension
Interaction Level
License/Openness
Description
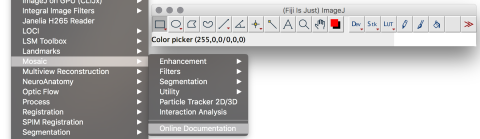
Image-processing algorithms developed at the MOSAIC Group for fluorescence microscopy. Tools included:
- 2D/3D single-particle tracking tool which can be used to track bright spots in 2D/3D movies over time.
- Optimal filament segmentation of 2D images.
- Curvature filters for image filtering, denoising, and restoration.
- Image naturalization for image enhancement based on gradient statistics of natural-scence images.
- Tool for automatically send and distribute jobs on clusters and get back the results.
- Multi-region image segmentation of 2D and 3D images without needing to know the number of regions beforehand.
- Squassh for globally optimal segmentation of piecewise constant regions in 2D and 3D images and for object-based co-localization analysis.
- Tool for inferring spatial interactions between patterns of objects in images or between coordinates read from a file.
- Tool for robust, histogram-based background subtraction well suited to correct for inhomogeneous illumination artifacts.
- A tool to estimate the Point-Spread Function of the microscopy out of 2D fluorescence images.
- A tool to measure the 3D Point-Spread Function of a confocal microscope from an image stack.
- Addition of synthetic Poisson-distributed noise to an image in order to simulate shot noise of various signal-to-noise ratios.
- Convolution of an image with a Bessel function in order to simulate imaging with a microscope.
- A utility to detect bright spots in images and estimate their center.
- A utility to create manual segmentations to be used as ground truth to test and benchmark automatic segmentation algorithms.
- A tool for replacing one color in an image with another color.
has function
has topic
Additional keywords
Entry Curator
Post date
10/06/2016 - 15:27
Last modified
05/08/2023 - 03:12