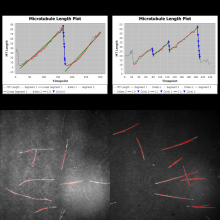
Description
MTrack is a tool, which detects, tracks, and measures the behavior of fluorescently labeled microtubules imaged by TIRF (total internal reflection fluorescence) microscopy. In such an in vitro reconstitution approach, stabilized, non-dynamic microtubule seeds serve as nucleation points for dynamically growing microtubules.
MTrack is a bi-modular tool. The first module detects and tracks the growing microtubule ends and creates trajectories. The second module uses these trajectories to fit models of dynamic behavior (polymerization and depolymerization velocities, catastrophe and rescue frequencies). It also computes statistics such as length and lifetime distributions when analyzing more than one movie (batch mode).