SuReSim
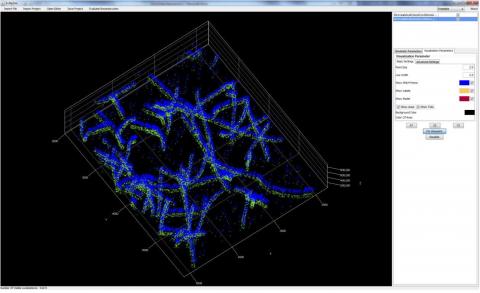
SuReSim (Super Resolution Simulation) is an open-source simulation software for Single Molecule Localization Microscopy (SMLM). The workflow of the SuReSim algorithm starts from a ground truth structure and lets the user choose to either directly simulate 3D localizations or to create simulated *.tiff-stacks that the user can analyze with any given SMLM reconstruction software. A 3D structure of any geometry, either taken from electron microscopy, designed de-novo from assumptions or known structural facts, is fluorophore-labeled in silico. A defined set of parameters is used to calculate and visualize the 3D localizations of the corresponding labels. The software package is accompanied with a library of model structures that can be imported and simulated. Users manual with tutorial provided.