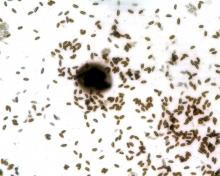
Mouse embryos
- Read more about Mouse embryos
- 198 views
Fluorescence microscopy cannot be used to image human embryos to determine embryo viability for in vitro fertilization because the introduction of exogenous fluorescent dyes is considered a toxic procedure. As a result, embryo viability has been measured primarily using differential interference contrast (DIC).